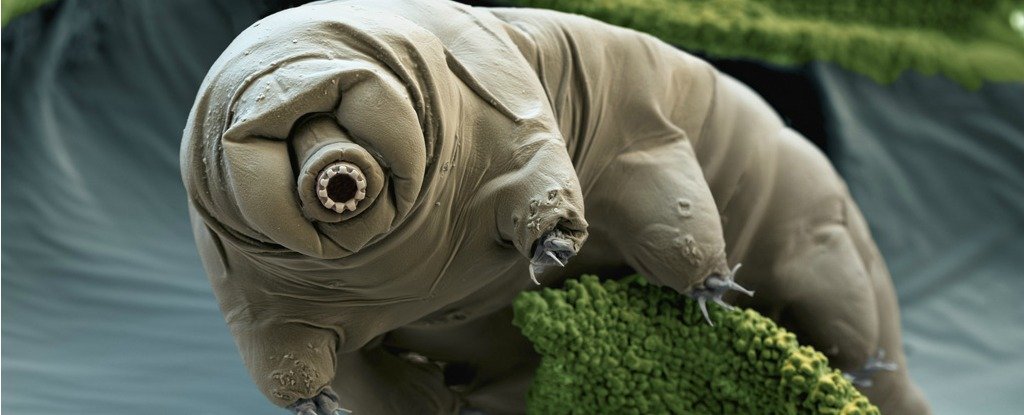
The reason the genome of the Tardigrade was such big news in November is that the group doing the bioinformatics analysis claimed that the genome contained 6,663 genes from bacteria, a full sixth of the genome, and twice as many horizontally transferred genes as have ever been seen in any other organism (Boothby et al, PNAS 112(52):15976-81. doi:
10.1073/pnas.1510461112. PMID: 26598659). This "weird science" observation was covered by National Geographic, Science News, Phys.org, Meta Science News, and of course the Univ. of North Carolina press site.
However, it seems quite clear now that this claim about horizontal DNA from bacteria (and maybe other phyla) in the genome of the Tardigrade was wrong. In fact, another group ( also working on the sequence of the exact same species has rapidly published a preprint manuscript on the bioRxiv preprint server "The genome of the tardigrade Hypsibius dujardini" that clearly refutes the claims of Boothby et al. and points out their mistakes in genome analysis: "Cross-comparison of the assemblies, using raw read and RNA-Seq data, confirmed that the overwhelming majority of the putative HGT candidates in the previous genome were predicted from scaffolds at very low coverage and were not transcribed."
It is quite easy to get contaminants when you are doing whole genome sequencing for a multicellular organism. You grind up your target species, extract DNA and put it into the sequencing machine. Any bacteria and other small organisms on the surface or in the gut come along for the ride and can contribute their DNA to the sequencing library. Surprisingly, a small amount of bacterial contaminating DNA (perhaps just 1%) can lead to a large number of bacterial contigs in the final genome assembly. I can think of a couple of reasons for this, based on the small size of bacterial genomes (~1 MB), vs metazoan genomes (most >100 MB). First, relative genome coverage of a contaminant bacteria will be much higher for each KB of sequence data, so the 1% of contaminating DNA may have deep coverage of a bacterial genome. Second, any two bacterial DNA fragments randomly selected from a library have a much higher chance to overlap (less complex genome), so they will assemble better.
There are a few QC steps that one can take on the raw data. There is a nice tool called Kraken (Wood DE, Salzberg SL Genome Biology 2014, 15:R46) that can quickly run through an entire FASTQ file (4 million reads per minute on a single core) and mark each read according to a set of reference genomes based on exact matching of 31 base k-mers. The Kraken team also make available a pre-built 4 GB database constructed from complete bacterial, archaeal, and viral genomes in RefSeq. DeconSeq is another good tool to find contaminants with an easy web interface. Of course, some legitimate reads from any target organism will share k-mer sized chunks with some bacteria, viruses, etc. (and some sequences from contaminating bacteria will not be in any database), so one has to make some tough choices about what to remove from the data before assembly.
After assembly, there are some additional steps one can take to flag contaminants. It is extremely helpful (I would now say required) to have some RNA-seq data from the same organism. RNA-seq data is prepared using a poly-A protocol, so no bacterial RNA contaminants should be present. Any contigs (with predicted genes) that do not contain a reasonable amount of aligned RNA-seq reads are highly suspect. Any contig that has predicted genes only from a different species is clearly a red flag.
While the authors of the original have not (yet) published a retraction, the citation in PubMed does carry a link to the refuting article provided by author Sujai Kumar
Rather than rant on about proper workflows for genome annotation (a best practices document does exist: Mark Yandell & Daniel Ence, Nature Reviews Genetics 13, 329-342 doi:10.1038/nrg3174) let me just say to the authors, the reviewers and the editors at PNAS that "EXTRODINARY CLAIMS REQUIRE EXTRODINARY EVIDENCE" (Carl Sagan). Or as said by Laplace: “The weight of evidence for an extraordinary claim must be proportioned to its strangeness.”